Günther R. Raidl
Institute of Computer Graphics
Vienna University of Technology
Karlsplatz 13/1861, 1040 Vienna, Austria
raidl@apm.tuwien.ac.at
Bryant A. Julstrom
Department of Computer Science
St. Cloud State University
St. Cloud, MN 56301 USA
julstrom@eeyore.stcloudstate.edu
The coding by which chromosomes represent candidate solutions is a
fundamental design choice in a genetic algorithm.
This paper describes a novel coding of spanning trees in a genetic algorithm
for the degree-constrained minimum spanning tree problem.
For a connected, weighted graph, this problem seeks to identify the shortest
spanning tree whose degree does not exceed an upper bound ![]() .
In the coding, chromosomes are strings of numerical weights associated with
the target graph's vertices.
The weights temporarily bias the graph's edge costs, and an extension
of Prim's algorithm, applied to the biased costs, identifies the feasible
spanning tree a chromosome represents.
This decoding algorithm enforces the degree constraint, so that all
chromosomes represent valid solutions and there is no need to discard,
repair, or penalize invalid chromosomes.
On a set of hard graphs whose unconstrained minimum spanning trees are
of high degree, a genetic algorithm that uses this coding
identifies degree-constrained minimum spanning trees that are on average
shorter than those found by several competing algorithms.
.
In the coding, chromosomes are strings of numerical weights associated with
the target graph's vertices.
The weights temporarily bias the graph's edge costs, and an extension
of Prim's algorithm, applied to the biased costs, identifies the feasible
spanning tree a chromosome represents.
This decoding algorithm enforces the degree constraint, so that all
chromosomes represent valid solutions and there is no need to discard,
repair, or penalize invalid chromosomes.
On a set of hard graphs whose unconstrained minimum spanning trees are
of high degree, a genetic algorithm that uses this coding
identifies degree-constrained minimum spanning trees that are on average
shorter than those found by several competing algorithms.
Keywords: Degree-constrained minimum spanning trees, weighted coding, genetic algorithms.
The coding by which chromosomes represent candidate solutions to a problem instance is a fundamental design choice in building a genetic algorithm (GA). Generally, a GA's designer may choose from several codings, some more conducive to genetic search than others.
Consider a connected, undirected graph G whose edges are labeled with
numerical costs.
A degree-constrained spanning tree on G is a spanning tree on G
none of whose vertices has degree greater than ![]() .
Identifying a minimum-cost degree-constrained spanning tree is
a computationally difficult problem with applications to
the design of telecommunication networks and integrated circuits.
Because of its hardness, the problem is addressed by heuristic methods,
including genetic algorithms.
.
Identifying a minimum-cost degree-constrained spanning tree is
a computationally difficult problem with applications to
the design of telecommunication networks and integrated circuits.
Because of its hardness, the problem is addressed by heuristic methods,
including genetic algorithms.
This paper presents a novel coding for spanning trees in a GA that searches for a degree-constrained minimum spanning tree. In this coding, a chromosome is a string of weights associated with the target problem instance's vertices. These weights temporarily bias the graph's edge costs, and an extension of Prim's algorithm identifies a degree-constrained spanning tree using the biased costs; the chromosome's fitness is the cost of the spanning tree under the original, unbiased costs. The decoding algorithm enforces the degree constraint, so that all chromosomes represent valid solutions and there is no need for the GA to discard, repair, or penalize invalid chromosomes. On a set of hard graphs whose unconstrained minimum spanning trees are of high degree, a steady-state GA that uses the weighted coding identifies degree-constrained spanning trees that are on average shorter than those found by several competing algorithms.
The following sections of this paper describe the degree-constrained minimum spanning tree problem; some previous genetic codings of spanning trees; the weighted coding of spanning trees; the genetic algorithm that uses the weighted coding; and comparisons of the GA's performance with that of several other algorithms on some challenging problem instances.
Given a connected, undirected graph G with n vertices,
a spanning tree T is a subgraph of
G that connects all of G's vertices and contains no cycles.
When numerical costs ![]() are associated with each edge (i,j),
a minimum spanning tree (MST) is a
spanning tree on G with the smallest possible total edge cost
are associated with each edge (i,j),
a minimum spanning tree (MST) is a
spanning tree on G with the smallest possible total edge cost
![]() .
Two well known algorithms, due to Prim [26] and Kruskal
[20], identify a MST of a connected, undirected graph
in polynomial time.
.
Two well known algorithms, due to Prim [26] and Kruskal
[20], identify a MST of a connected, undirected graph
in polynomial time.
The degree of a vertex is the number of edges in which it participates,
and the degree of a graph is the maximum degree of its vertices.
A variation of the MST problem bounds the degree of the spanning tree with
a constant ![]() ;
it seeks a spanning tree of minimum cost and degree no more than k:
a k-MST.
A 2-MST is a Hamiltonian path of minimum length.
Finding such a path is related to the familiar traveling salesman problem
and is NP-hard [12].
;
it seeks a spanning tree of minimum cost and degree no more than k:
a k-MST.
A 2-MST is a Hamiltonian path of minimum length.
Finding such a path is related to the familiar traveling salesman problem
and is NP-hard [12].
Often the vertices on which we seek MSTs are points in the plane, and edge costs are the Euclidean distances between these points. For this case, Monma and Suri [21] showed that there always exists a MST with degree no more than five. Papadimitriou and Vazirani [24] proved that finding a k-MST in the Euclidean plane is NP-hard when k = 3, and conjectured that it remains NP-hard when k = 4. More generally, the costs associated with the graph's edges are arbitrary; that is, we do not think of the vertices as residing in the plane, and the costs between them need not satisfy the triangle inequality. In this case, a minimum spanning tree on n points may have degree up to n - 1.
While polynomial-time heuristics exist for finding k-MSTs in the plane [11, 31], they are less effective on the general problem. One simple but effective heuristic is due to Narula and Ho [22]: They modified Prim's algorithm so that at each step it includes the cheapest eligible edge--one connecting a vertex currently in the (partial) spanning tree with one not yet connected--that does not violate the degree constraint. We refer to this heuristic as k-Prim.
A deceptively appealing coding of spanning trees for genetic search is
based on Prüfer's proof of Cayley's Formula, which identifies the number
of distinct spanning trees on a complete graph of n vertices as ![]() [7], [10, pp. 103-104,].
The proof establishes a one-to-one correspondence between spanning trees
on n vertices and strings of length n-2 over an alphabet of n symbols
by describing algorithms that derive a tree from its string and
vice versa [10, pp. 104-106,].
[7], [10, pp. 103-104,].
The proof establishes a one-to-one correspondence between spanning trees
on n vertices and strings of length n-2 over an alphabet of n symbols
by describing algorithms that derive a tree from its string and
vice versa [10, pp. 104-106,].
Despite its elegance, the Prüfer coding is problematic in GAs. As several observers have pointed out, symbols' meanings depend on their predecessors. Patterns of symbols do not represent consistent substructures of spanning trees, so that crossover may generate offspring whose trees do not resemble the trees of their parents, and the mutation of even one symbol may change the represented tree radically [23, 30].
Nonetheless, several researchers have used the Prüfer coding to represent candidate solutions in GAs for spanning tree problems. For example, Abuali, Schoenefeld, and Wainwright [1] used the Prüfer coding in a GA for the probabilistic minimum spanning tree problem. Operators included circular left shift, swap of two random positions, and random modification of a symbol. Zhou and Gen [32] presented a Prüfer-coding-based GA for the k-MST problem. This algorithm applied conventional genetic operators: uniform crossover and mutation by randomly modifying a symbol. Solutions that violated the degree constraint were repaired.
Kim and Gen [17] extended the Prüfer coding in a GA for a network design problem. A Prüfer string represented a spanning tree of service centers, and a second string indicated clusters of users the centers served. Again, chromosomes whose trees violated the problem's degree constraint were repaired. Gargano, Edelson, and Koval [13] extended the Prüfer coding with permutations in a GA for the time-dependent minimum spanning tree problem, in which edge costs depend on when the edges are included in the spanning tree; the Prüfer string represented a spanning tree, and the permutation the order in which vertices, and thus edges, were added to the tree.
Other projects have used more general codings whose represented structures
can include spanning trees, such as adjacency matrices [14],
indexes into lists of edges [8], and edge codings in which
when the value of the ![]() symbol
is j, the represented tree contains the edge (i,j) [3].
symbol
is j, the represented tree contains the edge (i,j) [3].
Knowles and Corne [18] described a GA for the k-MST problem. In their algorithm, chromosomes are sequences of integer values that influence the order in which k-Prim connects vertices to the growing spanning tree. Experiments on large, misleading graphs indicated the superiority of this approach to a dual simplex heuristic proposed by Boldon et al. [4]. Section 6 below compares our GA with both the algorithm of Corne and Knowles and the heuristic of Boldon et al.

Figure 1: (a) A set of points in the plane and their
coordinates; (b) a minimum cost spanning tree (of length 280.78) on
the points; and (c) the spanning tree of degree 3 on the points
corresponding to the chromosome (1.3, 0.4, 1.8, 1.5, 0.9, 0.8, 1.6,
2.1, 0.7, 0.5, 0.6, 1.8, 7); it has length 359.44. The chromosome's
last entry indicates that k-Prim began building the tree at vertex 7.
The coding the present GA employs is based on an ingenious coding of spanning
trees described by Palmer and Kershenbaum [23].
They encoded spanning trees as strings of numerical weights associated
with the graph's vertices.
To identify the tree such a chromosome represents, each weight ![]() is temporarily added to the costs of every edge in which vertex i
participates:
is temporarily added to the costs of every edge in which vertex i
participates:
![]()
Prim's algorithm then identifies a spanning tree from the biased costs. The chromosome's fitness is the cost of the tree using the original costs. Initial chromosomes consist of weights chosen at random from a uniform distribution. Mutation randomly resets a weight, and traditional crossover operators like uniform crossover can recombine chromosomes.
Palmer and Kershenbaum found that a weight-coded GA outperformed a good heuristic on the optimal communications spanning tree problem, and Abuali, Schoenefeld, and Wainwright [2] obtained better results with both a weighted coding and an edge coding than with the Prüfer coding in a GA for the probabilistic minimum cost spanning tree problem.
More generally, researchers have used weighted codings in GAs for a variety of combinatorial problems. These include the rectilinear Steiner problem [15], the shortest common supersequence problem [5], the 3-satisfiability problem [9], the traveling salesman problem [16], and the multiple container packing problem [27]. In each of these, chromosomes are strings of weights that temporarily bias parameters of the problem instance. A decoding algorithm identifies the structure a chromosome represents using the biased parameters, and the chromosome's fitness is that structure's fitness under the original parameters. Searching the space of weights implements a search of the target problem's search space.
One of us investigated biasing techniques in a weight-coded GA for the multiconstraint knapsack problem [28]. He found that multiplying parameters by weights selected from a log-normal distribution led to better solutions more quickly than did adding weights chosen from a uniform distribution.
We adopt the multiplicative scheme here.
Weights are chosen from the distribution ![]() ,
where
,
where ![]() is the normal distribution with mean 0 and
variance 1, and
is the normal distribution with mean 0 and
variance 1, and ![]() is a parameter called the biasing strength.
Edge costs are biased multiplicatively by these weights:
is a parameter called the biasing strength.
Edge costs are biased multiplicatively by these weights:
![]()
The k-Prim heuristic, applied to the biased edge costs, identifies the spanning tree a chromosome of such weights represents.
k-Prim grows a spanning tree from a start vertex, and in general it will not identify the same tree from different start vertices. The start vertex is particularly influential when a graph's underlying MST contains vertices of high degree. Clearly, fixing the start vertex might restrict the GA's search to poorer regions of the search space, so the coding is extended with an integer that represents the start vertex. The GA then adapts this choice along with the weights.
For example, Figure 1(a) depicts a set of 12 points in the plane and lists their coordinates. Figure 1(b) shows an unconstrained MST on these points; it has degree 5 and cost 280.78. A weighted chromosome that represents a 3-MST on these points consists of 12 float values and an integer, say
![]()
Vertex 1 has weight 1.3, vertex 2 has weight 0.4, and so on. The distance between vertices 1 and 2 is
![]()
so the biased distance between them is
![]()
The chromosome's last entry specifies vertex 7 as the start vertex. Figure 1(c) shows the tree k-Prim (with k = 3) identifies from the chromosome; using unbiased costs, it has cost 359.44.
Because k-Prim always identifies a spanning tree with degree no more than k, all chromosomes represent valid solutions, and there is no need for the GA to implement any of the usual constraint-handling strategies: discarding, repairing, or penalizing invalid chromosomes.
The weighted coding of spanning trees was implemented in an otherwise conventional steady-state GA. The algorithm selects chromosomes to be parents in tournaments of size three, and generates offspring from them via uniform crossover and a mutation that resets each gene to a new random value with a small probability (position-by-position mutation). Each new chromosome replaces the population's current worst chromosome, with one exception: To preserve phenotypic diversity, the GA discards any new chromosome that encodes a spanning tree already represented in the population [29].
The GA initializes its population with randomly generated chromosomes, except for one in which all the weights are 1.0 and only the starting vertex is selected at random. This chromosome represents the unbiased case and decodes to a spanning tree that k-Prim would find on its own. Early experiments confirmed that seeding the population in this way dramatically speeds the optimization process. Other preliminary experiments seeded the population with several chromosomes whose weights were all 1.0 but whose start vertices differed. This technique reduced the population's diversity and led to inferior results.
In the tests the following section describes,
![]() was set to 1.5, the GA's population held 100 chromosomes,
and the probability of mutating any one weight was 2/n, where n
is the number of vertices in the target graph.
was set to 1.5, the GA's population held 100 chromosomes,
and the probability of mutating any one weight was 2/n, where n
is the number of vertices in the target graph.
Boldon et al. [4] observed that when a large matrix of edge-costs is generated at random, the degree of a MST on the resulting graph rarely exceeds 3 or 4. They constructed more demanding problem instances from MSTs of high degree. Knowles and Corne [18] applied this approach to generate two sets of challenging k-MST instances, which we adopted to test the weight-coded GA.
The first problem set contains nine complete graphs of 50 to 200 vertices. Their unconstrained MSTs have degree 14 or 15. k-MSTs of maximum degree k = 4 were sought with k-Prim alone, the dual simplex method of Boldon et al., Knowles and Corne's GA, and the weight-coded GA. Both genetic algorithms were run 20 times through 10,000 evaluations on each instance; the deterministic heuristics were, of course, each applied once.

Table 1: 4-MST results for 9 randomly generated graphs
(![]() ). k-Prim indicates the
k-Prim heuristic alone; DS the dual simplex method of Boldon
et al.; K-GA the genetic algorithm of
Knowles and Corne; and W-GA the weight-coded GA.
). k-Prim indicates the
k-Prim heuristic alone; DS the dual simplex method of Boldon
et al.; K-GA the genetic algorithm of
Knowles and Corne; and W-GA the weight-coded GA.
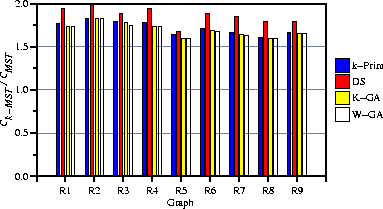
Figure 2: Average 4-MST results for the 9 randomly generated graphs
(![]() ).
).
Table 1 summarizes the results of these trials, reported as ratios of the lengths of shortest 4-MSTs to the lengths of unconstrained MSTs. Figure 2 shows these average results graphically. The two GAs performed almost identically, slightly better than k-Prim alone and the dual simplex method.
The second problem set consists of nine graphs made more difficult by the inclusion of edge weights intended to mislead heuristics. Again the graphs contain from 50 to 200 vertices; their unconstrained MSTs have degrees from 9 to 13. The algorithms sought k-MSTs with k = 5.
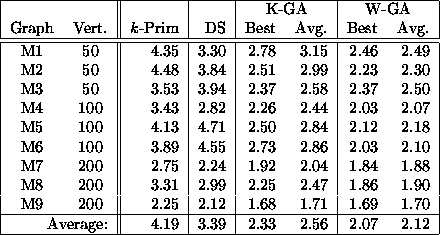
Table 2: 5-MST results for 9 misleading random graphs
(![]() ). Again, k-Prim indicates the
k-Prim heuristic alone; DS the dual simplex method of Boldon
et al.; K-GA the genetic algorithm of
Knowles and Corne; and W-GA the weight-coded GA.
). Again, k-Prim indicates the
k-Prim heuristic alone; DS the dual simplex method of Boldon
et al.; K-GA the genetic algorithm of
Knowles and Corne; and W-GA the weight-coded GA.
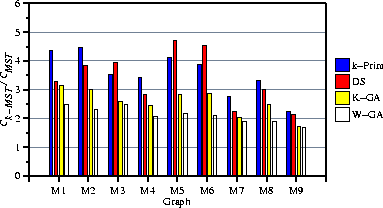
Figure 3: Average 5-MST results for the 9 misleading random graphs
(![]() ).
).
Table 2 and Fig. 3
summarize the results of these trials.
On these graphs, k-Prim performed poorly, the dual simplex method better,
and the two GAs best.
On average, the weight-coded GA identified significantly shorter trees
than did Knowles and Corne's GA, indicating that the weight-coded GA more
often escapes from local minima; the relative costs
![]() are 17% smaller
for the weight-coded approach.
Further, the best solutions of the weight-coded GA are usually
better and, except for M9, never worse then the corresponding best solutions of
Knowles and Corne's GA.
are 17% smaller
for the weight-coded approach.
Further, the best solutions of the weight-coded GA are usually
better and, except for M9, never worse then the corresponding best solutions of
Knowles and Corne's GA.
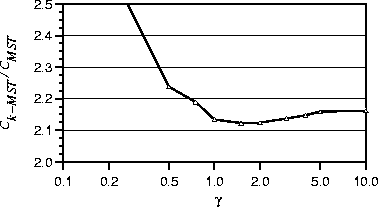
Figure 4: The effect of the biasing strength ![]() on the weight-coded
GA's average performance.
on the weight-coded
GA's average performance.
Figure 4 shows how the biasing strength ![]() affects the
quality of the weight-coded GA's solutions.
Each value in the graph was determined by averaging the results of 20
runs on each of the misleading problems M1 to M9.
For values of
affects the
quality of the weight-coded GA's solutions.
Each value in the graph was determined by averaging the results of 20
runs on each of the misleading problems M1 to M9.
For values of ![]() less than 0.5, the weighted coding is not
able to represent solutions significantly different from those k-Prim
would find on its own.
With larger
less than 0.5, the weighted coding is not
able to represent solutions significantly different from those k-Prim
would find on its own.
With larger ![]() , the GA can examine more of the problem's search space.
As
, the GA can examine more of the problem's search space.
As ![]() increases above 2.0, the solution quality again decreases;
the search becomes more diffuse.
Choosing
increases above 2.0, the solution quality again decreases;
the search becomes more diffuse.
Choosing ![]() yields the best results on average, and
in general it is better that
yields the best results on average, and
in general it is better that ![]() be too large than too small.
be too large than too small.
The weight-coded GA was also applied to the 9-vertex graph used by Zhou and Gen [32]. Their Prüfer-coded GA found an optimal solution 67% of the time in 25000 evaluations. In contrast, this problem turned out to be trivial for the weight-coded GA, which usually found an optimal solution in the initial population.
This paper has described the k-MST problem and a steady-state GA that encodes candidate solutions to the problem in a novel way. Each chromosome is a string of weights associated with the vertices of the target graph. These weights temporarily bias the graph's edge costs, and a modification of Prim's algorithm identifies the degree-k spanning tree a chromosome represents; this tree is evaluated using the original edge costs. Each chromosome also encodes a starting vertex for the decoding algorithm. In tests on hard and misleading k-MST instances, the weight-coded GA found degree-constrained spanning trees that are on average shorter than those identified by other heuristics.
More generally, this investigation and others illustrate the utility of weighted codings in GAs for combinatorial problems, particularly those with constraints. When, as here, the decoding algorithm enforces constraints, all chromosomes represent valid solutions, and there is no need to discard, repair, or penalize invalid chromosomes.