Copyright ACM, 2000
EEGS
OLVER - BRAIN ACTIVITY AND GENETIC ALGORITHMS
|
Paulo Aguiar |
André David |
Sandra Paulo |
Agostinho Rosa |
LaSEEB, ISR, Instituto Superior Técnico
- Av. Rovisco Pais, 1000 Lisboa, Portugal; l42585@alfa.ist.utl.pt
Keywords - EEG, Inverse Problem Solving, Genetic Algorithm, Brain Activity, Epilepsy.
Abstract
Genetic Algorithms (GAs) are a family of computational models inspired by evolution. This work tries to figure out what contribute could this class of algorithms give in an inverse problem solving frame: having an electroencephalogram (EEG), finding out where the spots of activity in the brain are. Our Brain is a mysterious machine of thoughts. As you read this text, you have billions of neurons working and creating complex, yet specific, electric potential patterns. We think the followed approach, in addition to the existing advanced analysis strategies, can provide an easily matchable piece to the big puzzle.
Introduction
- The Human Electroencephalogram
Every moment our brain cells generate millions of nervous impulses (nerve action potentials) and graded potentials (excitatory and inhibitory postsynaptic potentials) in individual neurons. These electrical potentials added together are called brain waves and reflect electrical activity of the cerebral cortex. Brain waves pass through the skull and can be detected by sensors called electrodes. A record of such waves is named electroencephalogram (EEG).
The human EEG was discovered by Berger (1929), when using a primitive galvanometer with a surface electrode placed on his son’s scalp, he recorded a rhythmic pattern of electrical oscillation. This signal was the electrophysiological response of cortical brain cells. It is now believed that the electrical potentials recorded in an EEG are produced in the pyramidal cell layer.
Many pyramidal cells and their dendrites are arranged vertically. The electrical activity of the brain can be explained by changes in membrane polarisation, which in turn cause the action potentials. The electrical potential is conducted through brain tissue, enter the membranes surrounding the brain, CSF (cerebrospinal fluid), continue on up through the skull to appear finally at the scalp, as depicted on figure 1.
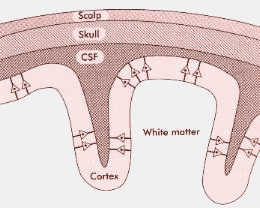
Figure 1 – Section of the head
At this point the potential is reduced from the milliVolt range (the membrane potential gradients and action potentials) to just a few microVolt.
The EEG is used to diagnose a large span of diseases such as epilepsy and other seizure disorders, tumours, brain trauma, hematomas and metabolic abnormalities, among others. Studies during periods of unconsciousness and confusion are also performed. The EEG may also be one criterion in establishing brain death as complete absence of brain waves in two EEGs taken 24 hours apart.
Epilepsy is the most common neurological disease: about 0.5 to 1% of the population suffers from it.
Typical EEG values are in the 20-100m V range. For EEGs with lesser values being recorded in averaged evoked potentials perhaps 10m V. Larger values are recorded in epilepsy and other disorders.
- Recording Techniques
An EEG is a record of the fluctuations of the electrical activity of large ensembles of neurons in the brain while the patient is sitting quietly or sleeping.
Neuroelectric signals are recorded as potential differences between electrodes. The recorded signals therefore depend intimately on the positions of the individual electrodes and how they are paired. Most laboratories performing routine clinical evaluations use the international 10-20 system of electrode placement. This system is based on 10% or 20% proportional distances between anatomic landmarks on the skull and head. The Nasion is located at the bridge of the nose immediately beneath the forehead, and the Inion is a bony protrusion located in the middle of the back of the head. The standard recording array for adults consists of 21 electrodes plus a ground electrode, as seen in figure 2.
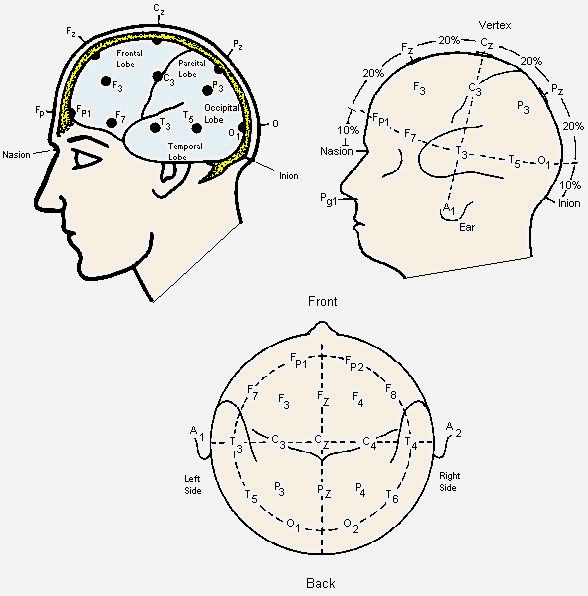
Figure 2 – Electrode positions in the 10-20 system
Each electrode is specified by a letter name related to the general underlying cortical region (Frontopolar - Fp; Frontal - F; Temporal - T; Occipital - O; Parietal - P) and a subscript reflecting its position relative to the midline. Even and odd numbers refer to the right and to the left hemisphere respectively, and the z refers to an electrode placed on the midline.
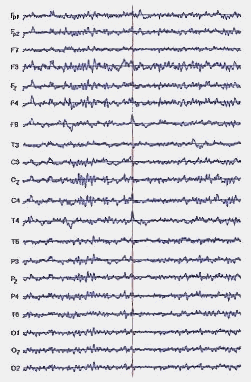
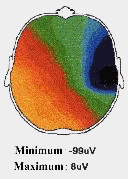
An EEG is a measure of the extracellular current flow associated with the summed activity of many individual neurons. The surface recorded potentials reflect predominantly the activity of cortical neurons in the area underlying the EEG electrode. Figure 3 illustrates a case of epilepsy.
Figure 3 – EEG example for epilepsy (from [12])
- Goals
The main objective of this work was not to solve an EEG with all the real conditions, such as tridimensional geometry for the brain, noise effects in the dipoles and other details.
Instead, we wanted to test the Genetic Algorithms under severe computational conditions and see what contributes they could give to this important problem. The EEG solving inverse problem a hard computational problem, with several biological and physiological constraints, and presently there is no program that gives satisfactory solutions in a big enough range of conditions in a short time.
Nowadays physiologists combine these different methods with different properties trying to improve results [9][12].
As we will see further ahead, the GA is able to take into account several constraints without evoking much more computational effort. It is in that field that we think GAs can give good contributions and can take a place in finding better solutions.
Biological Description
- Cellular Mechanisms
Having recorded the potentials, we can mathematically model where they were generated. This is called the inverse problem: the problem of finding system configuration from the final result. Mathematically there are an infinite number of initial source configurations that can generate a given scalp potential. Empirical and theoretical evidence suggests that the majority of the observed electromagnetic signals arise from the cortical gray matter, specifically the apical dendrites of the cortical pyramidal cells. These neurons have a columnar organization, oriented perpendicularly to the cortical sheet as figure 1 suggests. Thus, if the shape of the cortical sheet is known, this information can be used to constrain the locations and orientations of the cortical sources.
EEG is based on the theory of volume conduction, which describes the flow of ionic current generated by nerve cells through the extracellular space as depicted in figure 4.
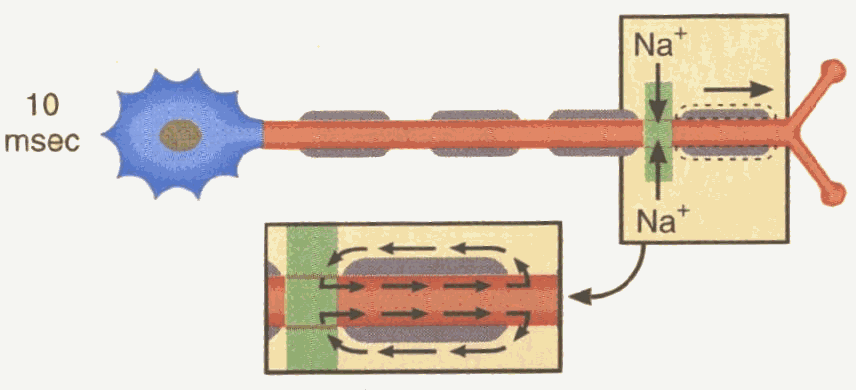
Figure 4 – A neuron
The sum of the ionic currents of many thousands of neurons located under the recording electrode generates potential changes, recorded at the scalp. The net ionic current is recorded as a voltage across the resistance of the extracellular space.
The extracellular current sinks and sources (and associated volume currents) cause an electrical difference of potential in the conductive media, with the magnitude of the potential decreasing with increasing distance from neuron. See figure 5.
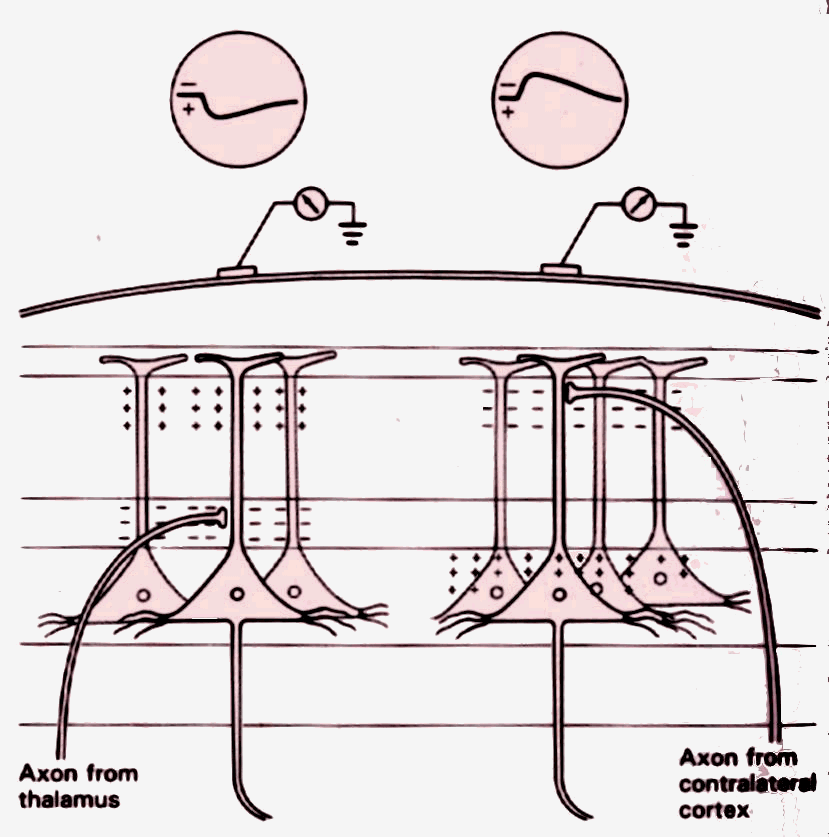
Figure 5 – Cortical neurons
The potential behaves as if it were produced by a current flow between two poles. This dipole approximation can be accurately applied to the potential pattern generated by focal population of neurons, such as those that make up a cortical column. Notice that the potential pattern generated by complex activation of the thousands of neurons comprised in a cortical column is, at a measuring distance of a centimetre or more, mostly dipolar. The potential produced by a dipole, in a homogenous surrounding medium, is proportional to the inverse square of the distance, a 1/r2 law.
The electrical potential difference recorded between two electrodes reflect weighted contributions from all current sources within the medium, so strong activity at a long distance will be slightly influential. The recorded signal comes mainly from neurons near the tip of the electrode and only to a small extent from more distant neurons.
The specification of the potential pattern, which a particular current configuration generates within a homogenous conductive medium, is relatively straightforward as given by Maxwell’s equations. However, the human head is not a homogenous volume. There are local inhomogeneities within the brain and significant conductivity barriers between the brain, CSF, skull and scalp.
GAs Meet EEGs
- Introduction
Given knowledge of the electrical properties and geometry of the head, the surface potential pattern generated by a particular pattern of intracranial neuron currents is specified by Maxwell’s equations. However the current pattern cannot be specified uniquely because currents elements may produce surface potentials that cancel each other. This inverse problem of specifying generative neuronal activity is ill posed and unsolvable in its most general, unrestricted form. Although, by making assumptions about the conductive properties of the head and the configuration of the generative currents, iterative procedures can be used to determine best-fitting (in the least squares or another norm sense) source configurations [12]. In the analysis of neuroelectric data, the head is modelled as a spherical volume conductor of multiple concentric shells of differing but homogenous electrical conductivity. A four-shell model is common with the shells representing the brain, CSF, skull, and scalp. Currents associated with the neuronal activity of a focal population are mathematically modelled as if they were generated by a dipole source. Provided that the measured potential pattern mostly reflects activity from a single focal neuronal population, and provided that the distance between scalp surface recording sites and the location of the neuronal activity is large comparing to the spatial extent of the activated neuronal region, the dipole approximation is quite reasonable.
Given these assumptions, mathematical procedures can be used to derive the position, orientation, and strength of the dipole that best accounts (in a least squares or other norm sense) for the measured potential pattern.
- GAs
Genetic algorithms can be a powerful tool solving problems and simulating natural systems in a variety of scientific fields.
Why use evolution as an inspiration for solving computational problems? The mechanisms of evolution seem well suited for some of the most pressing computational problems in many fields. Many computational problems require searching through a huge number of possible solutions. In evolutionary computation the rules are typically natural selection with variation due to crossover and/or mutation. Biological evolution is a method of searching among an enormous number of possible solutions.
At this point it is useful to formally introduce some of the biological terminology that we used throughout our simulation. These terms are used in the spirit of the analogy with real biology.
All living organisms are made up of cells, each one containing one or more chromosomes, strings of DNA, that serves to identify the organism. A chromosome can be conceptually divided into genes, which roughly can be thought as a codification for a property, like eye colour. The different eye colours, like blue or brown, are called alleles.
Each gene is located at a particular position on the chromosome. The complete collection of chromosomes of an organism is called its genome. The genotype refers to the particular set of genes contained in a genome. The physical and mental characteristics, such as height or intelligence, give rise to the organism’s phenotype. Organisms whose chromosomes are arrayed in pairs are called diploid and whose chromosomes are unpaired, haploid.
Another important concept is the fitness of an organism. It is typically defined as the probability that the organism will live to reproduce (viability) or as a function of the number of offspring the organism has (fertility).
GAs work by discovering, emphasising, and recombining good building blocks of solutions in a highly parallel fashion. This notion of building blocks can be formalised as schemata, and corresponds to constellations of genes that work together to affect some adaptation in an organism.
Given a particular potential distribution, how do we know if a GA is good method to use? There is no rigorous answer!
- Method Implementation
Our organisms were defined as dipole configurations. They were set as haploid: one chromosome has all the information about all the dipoles. To each dipole corresponds a gene, with information of it’s the position, magnitude and orientation.
A very important matter is that this choice of genes has infinite alleles: all the parameters are real numbers so they can be set as any value. Note that the genotype of our species is the same as the phenotype.
When we speak of methods for EEG solving we mean mathematical structures that match the external read potentials with dipoles in brain locations, and with electrical features, that have physiological meaning.
In terms of Maxwell equations, this problem has infinite solutions so strict biological/physical conditions/constraints are needed to get reasonable results.
Two kinds of methods are implemented in the existing computational programs in finding such dipoles:
- Let a very small and variable number of dipoles be free in their locations inside de brain;
- Set a large matrix with fixed dipoles, and vary only their magnitude and orientation.
To fulfil our purposes, we decided for the second method, which implies a larger computational effort. Some present programs use a matrix with about 250 dipoles. They use a method that requires matrix inversion and where minimising a linear or quadratic norm chooses the solution (L1/L2 estimation methods [5][10]). This method is of common use in the EEG centres and takes about ten minutes to give a solution, which, sometimes is not reasonable in physiological terms.
In this present study we worked with the Fp1, Fp2, F8, T4, T6, O2, O1, T5, T3 and F7 electrodes, 10 in total. All together, they draw a line placed in a plane, cf. figure 2. It was inside this 2D-closed line that we tried to find solutions for the dipole distribution.
Using the common circular approximation for this brain section we used a 600-dipole matrix where the dipoles were placed according to the following criteria: like the surface recorded potentials reflect predominantly the activity of cortical neurons we only distribute dipoles in this particular area. This gives us a particular distribution where the white matter has no dipoles (medium radius of 40mm). In the cortex zone, the dipoles are placed, so their density gets greater until the peripheral surface. Note that the ratio of cortex/white matter has a non-linear growth with increasing radius (radius from 40mm to 70mm).
|
Figure 6 – 2D brain section |
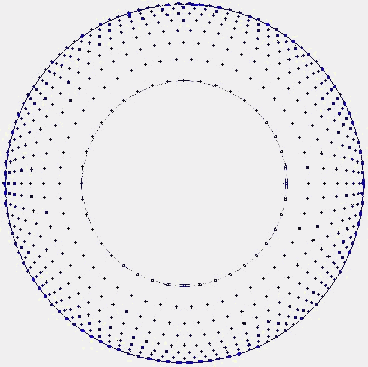
Figure 7 – The 600 dipole distribution |
This is an empirical disposition, but is already a light physiological constraint with better results than a homogeneous distribution, as tested.
By now, our organisms are well defined: one chromosome has 600 genes where each gene encodes the dipole location inside de brain section - and two genetic free parameters: magnitude and orientation. So, roughly, we will have a species that has 1200 genetic parameters with infinite alleles.
The next step in a GA after deciding codification, is how to perform natural selection. By this we mean what criteria we use to choose the individuals in the population that will create offspring for the next generation and how many in the offspring will be created. Another important thing is to decide what kind of external conditions can alter the genotype of the individuals.
The purpose of selection is, of course, to emphasise the fittest individuals in the population hoping that their offspring will in turn have even higher fitness by trading genetic material with others. Selection has to be balanced with variation from crossover and mutation: too-strong selection may imply suboptimal highly fit individuals taking over the population, reducing the diversity needed for further range and progress; too-weak selection may result in too-slow evolution.
Genetic Operators
A large use of the schemata theory was used to perform reproduction.
Constellations of genes were set and grouped to perform areas of influence for each electrode. The circumference of the brain section was divided into ten sectors 36º each, in which the electrodes were placed at the middle (18º) and at a radius of the scalp.
The potential is proportional to 1/r2 where r is the distance between the dipole and the point where we read the potential. So, as an example, dipoles that create a certain potential in the F8 electrode don’t have much influence in the O2.
The reproduction between two individuals represents, in this approach, the exchange of those sections. In terms of chromosomes, reproduction implies that the two chromosomes brake into ten pieces each and recombine after exchanging genetic material. This crossover, in a ten times version, give us some innovation and variation but it is not enough to guarantee that the population will not fall in some particular locus with permanent fixation in the search space. Mutation, hence, plays a very important role.
In our program we defined to kinds of mutation:
- Mutants, generated with slight random variation of their genetic code - this help us to solve the electrical problem which have infinite solutions;
- Genetic viruses, sent sometimes to infect the organisms with non-reasonable biological/physiological dipole configurations.
It is in this last point that we think GA can make a difference: we can make as much genetic viruses as we want to satisfy bio-constraints such as magnitude, influence and so on.
The work basis was the canonical GA. In terms off flow chart:
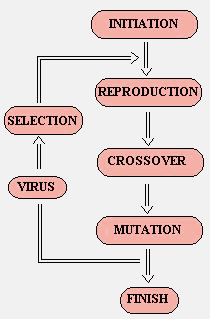
Figure 8 – The implemented GA flowchart
The most important thing in GAs, is although, the selection. Our fitness function was developed over a 10 element vector, where each element represents the difference between the value read in a electrode and the potential value calculated by the dipole configuration. Each chromosome has this vector virtually attached to it. All the actions taken in the chromosome directly affect this vector, including the genetic viruses.
As a statistic value, the sum of the squares of the differences (S D 2) is taken to measure how fit, in an electrical way, the chromosome is. Using m V as the potential unit, a S D 2=1 means that each electrode has an average difference of 0.32m V between the real value and the calculated one. Remember that the common values for EEG readings are from 10 to 20 m V (in absolute value). Organisms with lesser S D 2 have more probability to reproduce.
The genetic virus only attacks individuals that do not satisfy some physiological criteria. Sometimes, the virus completely destroys the organism's genotype leading him to death.
In the environment created we set the following law of surviving (and then reproducing):
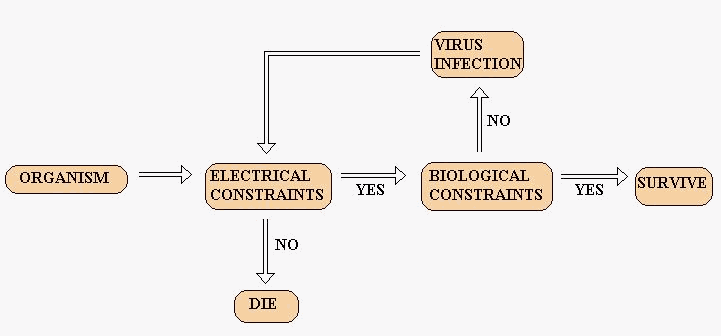
Figure 9 – Survival flowchart
To avoid losing precious organisms with special dipole configurations, we apply another operator, the elitism. In this way, we force the GA to retain some of the best individuals at each generation, comparing also with their parents. Only with the use of elitism one can assert that a GA can converge to the best solution – albeit the infinite time it can take.
- Results
Our program – the most obvious result of this work - can be tested at the Internet by anyone by going to
http://surf.to/EEGSolver. This binary has a fixed population of 30 organisms. In each reproduction 16 of them die. In terms of time, the GA program takes about 4 minutes, exempting time spent in kernel to server and server to client communications, to find a solution with S D 2<1. The current computer running the server and kernel, theta.ist.utl.pt, has 400MHz Alpha processors. Adding many bio-constraints in form of genetic viruses increases, obviously, the S D 2<1 time. The Internet version has only one virus acting which enforces that mutual influence between neighbour dipoles cannot have magnitudes and orientations too different – a smoothing virus. The addition of this virus implied an increase of about one minute. More viruses can represent more accurate results, in biological terms, and bit more of S D 2<1 time. But the good thing is that there are no limits for the number of viruses.
Due to the fact that GAs invokes a lot of probability parameters, we can play with that to have different time efforts to obtain a solution. The 4 minutes represent a large effort on parameter optimisation.
All the benchmarks were made against theoretical cases; after setting a particular zone(s) of activity in a virtual 2D-brain section and calculating the potentials at the 10 electrodes, those values were sent to EEGSolver to find, again, the original spots of activity.
Better results were found when the activity was at the peripheral brain surface. This arises form our dipole distribution.
The GA also shows to be useful in situations of more than one spot of activity, although for the virus we have used, these zones appeared blurred. Some of the most common programs don’t even detect the second spot of activity.
- Future Work
First of all… receive the 21 electrode potentials and find configurations in 3D-brain space. In terms of GA, the one thing that would change is the information in each gene and the calculation of the fitness: the potential function would now bear three coordinates. Everything else would remain basically the same in the GA.
Perhaps some effort could be taken in finding good viruses that could bring good results where the existing programs fail.
- Conclusions
GA may not substitute the others algorithms, but we believe it can be helpful. Depending on the implemented viruses, it can give better results in particular situations. But still the best way to study an EEG is combining several methods, and is clear that AG, with an EEGSolver type program, is important among them.
- Acknowledgements
We would like to thank Professor Ducla Soares and Doutora Carla Silva, both from the Instituto de Biofísica e Ciências Biomédicas at Lisbon for all their help and information about what is done in electroencephalography and what could be better in the computational solving programs. We which also to thank the Calouste Gulbenkian Foundation for the support.
- References
[
1] Haines, S., EEG , http://www.mhri.edu.au/bdl/papers.
[
2] Kandel, E., Schwartz, J., Jessell, T., 1991. Principles of Neural Science, pp. 95-122 and 777-791, 3rded. Prentice-Hall International Inc..
[
3] Kandel, E., Schwartz, J., Jessell, T., 1995. Essentials of Neural Science and Behaviour, pp. 161-177. Prentice-Hall International Inc..
[
4] Lopes da Silva, F.. Dynamics of EEGs as Signals of Neuronal Populations: Models and Theoretical Considerations, Chapter 4, pp. 63-77.
[
5] Maltez, J.C., Silva, C., Arriaga, A., Ducla-Soares, E., 1997. EEG Source Localization with L1 and L2 Minimum Norm Estimations – A Model Study, http://alf1.cii.fc.ul.pt/~arriaga/brain/sld001.htm, World Congress on Medical Physics and Biomedical Engineering, Nice, France,.
[
6] Mitchell, Melanie, 1998. An Introduction to Genetic Algorithms. MIT Press.
[
7] Oostendorp, T., Oosterom, A.,1989. Source Parameter Estimation in Inhomogeneous Volume Conductors of Arbitrary Shape, IEEE Transactions on Biomedical Engineering, Vol. 36, No. 3, March.
[
8] Orrison, Jr., Lewine, J., Sanders, J., Hartshorne, M., 1995. Functional Brain Imaging, pp.327-368. Mosby-Year Book,
[
9] Posner, M., Raichle, M., 1994. Images of Brain. Scientific American Library..
[
10] Reilly, Edward L.. EEG Recording and Operation of the Apparatus, pp.104-124.
[
11] Sarvas, J., Hämäläinen, M., 1989. Realistic Conductivity Geometry Model of the Human Head for Interpretation of Neuromagnetic Data. IEEE Transactions on Biomedical Engineering, Vol. 36, No. 2, February.
[
12] Silva, Carla, 1998. Processamento de Dados Electroencefalográficos - Aplicações à epilepsia (Ph.D. these). Faculdade de Ciências da Universidade de Lisboa.
[
13] Tortora, G., Grabowski, S., 1996. Principles of Anatomy and Physiology, 8th ed.. Harper Collins.
[
14] Whitley, D., A Genetic Algorithm Tutorial, http://www.geneticprogramming.com/Tutorial.
Copyright 2000 ACM
Permission to make digital or hard copies of all or part of this work for
personal or classroom use is granted without fee provided that copies are
not made or distributed for profit or commercial advantage and that copies
bear this notice and the full citation on the first page. To copy
otherwise, to republish, to post on servers or to redistribute to lists,
requires prior specific permission and or fee.
SAC 2000 March 19-21 Como, Italy
(c) 2000 ACM 1-58113-239-5/00/003>...>$5.00